Graph
Last updated: May 2, 2026Table of Contents
- Overview
- Key Properties
- Core Characteristics
- Graph Representations
- Problem Categories
- Category 1: Graph Traversal
- Category 2: Shortest Path
- Category 3: Union-Find (DSU)
- Category 4: Topological Sort
- Category 5: Bipartite Graphs
- Category 6: Minimum Spanning Tree
- Templates & Algorithms
- Template Comparison Table
- Universal Graph Template
- Template 1: BFS Traversal
- Template 2: DFS Traversal
- Template 3: Union-Find (DSU)
- Template 4: Topological Sort (DFS)
- Template 5: Topological Sort (BFS/Kahn’s)
- Template 6: Bipartite Graph Checking
- Template 7: Shortest Path Algorithms
- Template 8: Tarjan’s Algorithm (Graph Connectivity)
- Problems by Pattern
- Graph Traversal Problems
- Shortest Path Problems
- Union-Find Problems
- Topological Sort Problems
- Bipartite Problems
- Advanced Graph Problems
- 2) LC Example
- 2-1) Closest Leaf in a Binary Tree
- 2-2) Number of Connected Components in an Undirected Graph
- 2-3) Clone Graph
- 2-4) Bus Routes
- 2-5) Course Schedule
- Decision Framework
- Pattern Selection Strategy
- Algorithm Selection Guide
- Summary & Quick Reference
- Complexity Quick Reference
- Graph Building Patterns
- Common Patterns & Tricks
- Problem-Solving Steps
- Common Mistakes & Tips
- Interview Tips
- Related Topics
- 2-6) Find Eventual Safe States
Graph Algorithms
Overview
Graph algorithms are techniques for solving problems on graph data structures consisting of vertices (nodes) and edges (connections between nodes).
Key Properties
- Time Complexity: O(V + E) for traversal, varies for other algorithms
- Space Complexity: O(V) for adjacency list, O(V²) for matrix
- Core Idea: Model relationships and connections between entities
- When to Use: Network problems, dependencies, paths, connectivity
- Key Algorithms: BFS, DFS, Dijkstra, Union-Find, Topological Sort
Core Characteristics
- Directed vs Undirected: One-way or two-way edges
- Weighted vs Unweighted: Edges with or without costs
- Cyclic vs Acyclic: Contains cycles or not
- Connected vs Disconnected: All nodes reachable or not
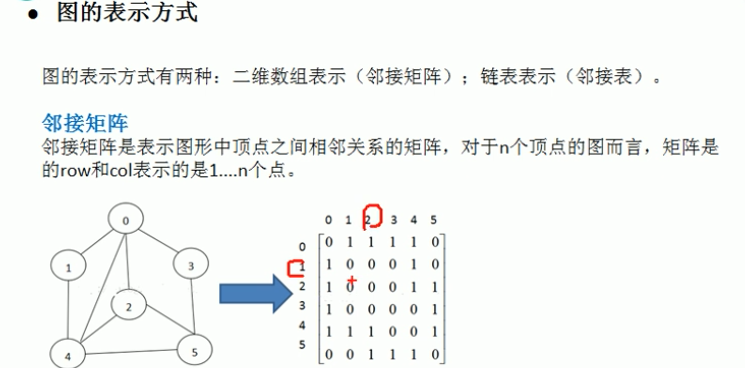
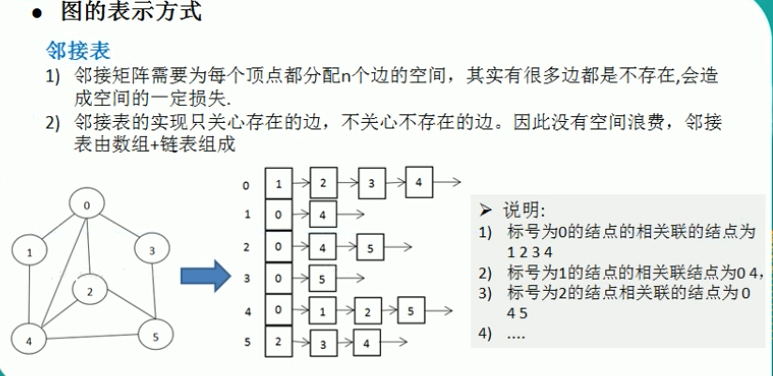
Graph Representations
- Adjacency List: Space O(V + E), efficient for sparse graphs
- Adjacency Matrix: Space O(V²), efficient for dense graphs
- Edge List: Space O(E), simple but less efficient
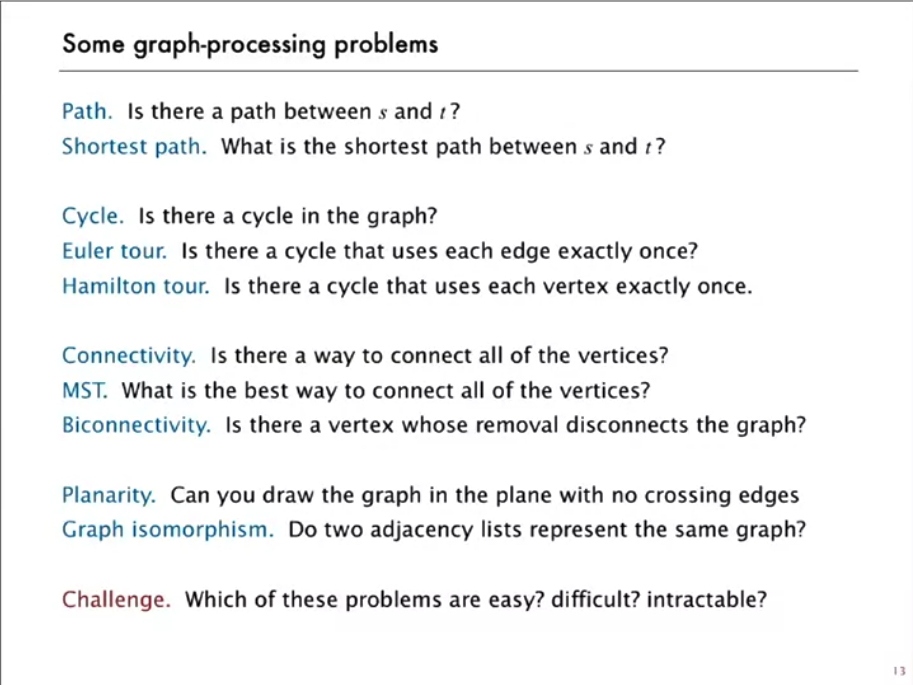

Problem Categories
Category 1: Graph Traversal
- Description: Explore all nodes using BFS or DFS
- Examples: LC 200 (Number of Islands), LC 133 (Clone Graph)
- Pattern: Visit all connected components
Category 2: Shortest Path
- Description: Find minimum distance between nodes
- Examples: LC 743 (Network Delay), LC 787 (Cheapest Flights)
- Pattern: Dijkstra, Bellman-Ford, Floyd-Warshall
Category 3: Union-Find (DSU)
- Description: Detect cycles, find connected components
- Examples: LC 684 (Redundant Connection), LC 721 (Accounts Merge)
- Pattern: Union by rank, path compression
Category 4: Topological Sort
- Description: Order nodes with dependencies
- Examples: LC 207 (Course Schedule), LC 210 (Course Schedule II)
- Pattern: DFS or Kahn’s algorithm (BFS)
Category 5: Bipartite Graphs
- Description: Check if graph can be colored with 2 colors
- Examples: LC 785 (Is Graph Bipartite), LC 886 (Possible Bipartition)
- Pattern: BFS/DFS with coloring
Category 6: Minimum Spanning Tree
- Description: Connect all nodes with minimum cost
- Examples: LC 1135 (Connecting Cities), LC 1584 (Min Cost Connect Points)
- Pattern: Kruskal’s or Prim’s algorithm
Templates & Algorithms
Template Comparison Table
| Template Type | Use Case | Time Complexity | When to Use |
|---|---|---|---|
| BFS | Level-order, shortest path | O(V + E) | Unweighted shortest path |
| DFS | All paths, cycle detection | O(V + E) | Explore all possibilities |
| Union-Find | Connected components | O(α(n)) | Dynamic connectivity |
| Dijkstra | Weighted shortest path | O((V+E)logV) | Non-negative weights |
| Topological | Dependencies | O(V + E) | DAG ordering |
Universal Graph Template
def graph_algorithm(n, edges):
# Build adjacency list
graph = collections.defaultdict(list)
for u, v in edges:
graph[u].append(v)
graph[v].append(u) # For undirected
# Track visited nodes
visited = set()
# Process each component
result = 0
for node in range(n):
if node not in visited:
# Process component
process_component(node, graph, visited)
result += 1
return result
Template 1: BFS Traversal
def bfs_template(graph, start):
"""Breadth-first search template"""
from collections import deque
visited = set([start])
queue = deque([start])
level = 0
while queue:
# Process level by level
size = len(queue)
for _ in range(size):
node = queue.popleft()
# Process node
for neighbor in graph[node]:
if neighbor not in visited:
visited.add(neighbor)
queue.append(neighbor)
level += 1
return level
Template 2: DFS Traversal
def dfs_template(graph, start):
"""Depth-first search template"""
visited = set()
path = []
def dfs(node):
visited.add(node)
path.append(node)
for neighbor in graph[node]:
if neighbor not in visited:
dfs(neighbor)
# Backtrack if needed
# path.pop()
dfs(start)
return visited
Template 3: Union-Find (DSU)
class UnionFind:
def __init__(self, n):
self.parent = list(range(n))
self.rank = [0] * n
self.count = n
def find(self, x):
"""Find with path compression"""
if self.parent[x] != x:
self.parent[x] = self.find(self.parent[x])
return self.parent[x]
def union(self, x, y):
"""Union by rank"""
px, py = self.find(x), self.find(y)
if px == py:
return False
if self.rank[px] < self.rank[py]:
px, py = py, px
self.parent[py] = px
if self.rank[px] == self.rank[py]:
self.rank[px] += 1
self.count -= 1
return True
def connected(self, x, y):
return self.find(x) == self.find(y)
Template 4: Topological Sort (DFS)
def topological_sort_dfs(n, edges):
"""Topological sort using DFS"""
graph = collections.defaultdict(list)
for u, v in edges:
graph[u].append(v)
# 0: unvisited, 1: visiting, 2: visited
state = [0] * n
result = []
def dfs(node):
if state[node] == 1: # Cycle detected
return False
if state[node] == 2: # Already processed
return True
state[node] = 1 # Mark as visiting
for neighbor in graph[node]:
if not dfs(neighbor):
return False
state[node] = 2 # Mark as visited
result.append(node)
return True
for i in range(n):
if state[i] == 0:
if not dfs(i):
return [] # Cycle exists
return result[::-1]
Template 5: Topological Sort (BFS/Kahn’s)
def topological_sort_bfs(n, edges):
"""Kahn's algorithm for topological sort"""
from collections import deque
graph = collections.defaultdict(list)
indegree = [0] * n
for u, v in edges:
graph[u].append(v)
indegree[v] += 1
queue = deque([i for i in range(n) if indegree[i] == 0])
result = []
while queue:
node = queue.popleft()
result.append(node)
for neighbor in graph[node]:
indegree[neighbor] -= 1
if indegree[neighbor] == 0:
queue.append(neighbor)
return result if len(result) == n else []
Template 6: Bipartite Graph Checking
Definition: A graph is bipartite if its vertices can be colored using only two colors such that no two adjacent vertices have the same color. Equivalent to checking if the graph has no odd-length cycles.
Time Complexity: O(V + E) - visit each vertex and edge once Space Complexity: O(V) - for color array and queue/recursion stack
Use Cases:
- Graph coloring problems
- Matching problems (assignment, scheduling)
- Conflict detection
- Resource allocation
- Network flow problems
Key Properties:
- All trees are bipartite
- Graphs with odd cycles are NOT bipartite
- Complete bipartite graphs K(m,n) are bipartite
- Can be solved using BFS or DFS with 2-coloring
Approach 1: BFS with Coloring
def is_bipartite_bfs(graph):
"""Check if graph is bipartite using BFS"""
from collections import deque
n = len(graph)
colors = [-1] * n # -1: uncolored, 0: color A, 1: color B
# Handle disconnected components
for start in range(n):
if colors[start] != -1:
continue
# BFS coloring
queue = deque([start])
colors[start] = 0
while queue:
node = queue.popleft()
for neighbor in graph[node]:
if colors[neighbor] == -1:
# Color with opposite color
colors[neighbor] = 1 - colors[node]
queue.append(neighbor)
elif colors[neighbor] == colors[node]:
# Same color conflict - not bipartite
return False
return True
Approach 2: DFS with Coloring
def is_bipartite_dfs(graph):
"""Check if graph is bipartite using DFS"""
n = len(graph)
colors = [-1] * n
def dfs(node, color):
colors[node] = color
for neighbor in graph[node]:
if colors[neighbor] == -1:
# Recursively color with opposite color
if not dfs(neighbor, 1 - color):
return False
elif colors[neighbor] == colors[node]:
# Same color conflict
return False
return True
# Check each component
for i in range(n):
if colors[i] == -1:
if not dfs(i, 0):
return False
return True
Approach 3: Union-Find (Advanced)
def is_bipartite_union_find(n, edges):
"""Check bipartite using Union-Find for conflict detection"""
class UnionFind:
def __init__(self, n):
self.parent = list(range(2 * n)) # 2n for opposite groups
def find(self, x):
if self.parent[x] != x:
self.parent[x] = self.find(self.parent[x])
return self.parent[x]
def union(self, x, y):
px, py = self.find(x), self.find(y)
if px != py:
self.parent[px] = py
def connected(self, x, y):
return self.find(x) == self.find(y)
uf = UnionFind(n)
# For each edge (u,v), union u with opposite of v, and v with opposite of u
for u, v in edges:
if uf.connected(u, v): # Same group conflict
return False
# u should be in opposite group of v
uf.union(u, v + n) # u with opposite of v
uf.union(v, u + n) # v with opposite of u
return True
Grid-based Bipartite Check
def is_bipartite_grid(grid):
"""Check if grid graph is bipartite (checkerboard pattern)"""
rows, cols = len(grid), len(grid[0])
colors = [[-1] * cols for _ in range(rows)]
directions = [(0, 1), (1, 0), (0, -1), (-1, 0)]
def bfs(start_r, start_c):
from collections import deque
queue = deque([(start_r, start_c)])
colors[start_r][start_c] = 0
while queue:
r, c = queue.popleft()
for dr, dc in directions:
nr, nc = r + dr, c + dc
if 0 <= nr < rows and 0 <= nc < cols and grid[nr][nc] == 1:
if colors[nr][nc] == -1:
colors[nr][nc] = 1 - colors[r][c]
queue.append((nr, nc))
elif colors[nr][nc] == colors[r][c]:
return False
return True
# Check each component
for i in range(rows):
for j in range(cols):
if grid[i][j] == 1 and colors[i][j] == -1:
if not bfs(i, j):
return False
return True
Enhanced Template with Partition Information
def bipartite_partition(graph):
"""
Returns bipartite partition if exists, None otherwise
Returns: (setA, setB) or None
"""
n = len(graph)
colors = [-1] * n
def dfs(node, color):
colors[node] = color
for neighbor in graph[node]:
if colors[neighbor] == -1:
if not dfs(neighbor, 1 - color):
return False
elif colors[neighbor] == colors[node]:
return False
return True
# Check bipartite property
for i in range(n):
if colors[i] == -1:
if not dfs(i, 0):
return None
# Create partition sets
setA = [i for i in range(n) if colors[i] == 0]
setB = [i for i in range(n) if colors[i] == 1]
return setA, setB
Related Problems & Examples
LC 785: Is Graph Bipartite
def isBipartite(self, graph):
"""LC 785 - Standard bipartite check"""
n = len(graph)
colors = {}
def dfs(node, color):
colors[node] = color
for neighbor in graph[node]:
if neighbor in colors:
if colors[neighbor] == colors[node]:
return False
else:
if not dfs(neighbor, 1 - color):
return False
return True
for i in range(n):
if i not in colors:
if not dfs(i, 0):
return False
return True
LC 886: Possible Bipartition
def possibleBipartition(self, n, dislikes):
"""LC 886 - Build graph from dislike relationships"""
from collections import defaultdict
# Build adjacency list from dislikes
graph = defaultdict(list)
for u, v in dislikes:
graph[u].append(v)
graph[v].append(u)
colors = {}
def dfs(node, color):
colors[node] = color
for neighbor in graph[node]:
if neighbor in colors:
if colors[neighbor] == colors[node]:
return False
else:
if not dfs(neighbor, 1 - color):
return False
return True
for i in range(1, n + 1):
if i not in colors:
if not dfs(i, 0):
return False
return True
Applications & Variations
1. Maximum Bipartite Matching
def max_bipartite_matching(graph, n, m):
"""Find maximum matching in bipartite graph"""
match = [-1] * m
def dfs(u, visited):
for v in graph[u]:
if not visited[v]:
visited[v] = True
if match[v] == -1 or dfs(match[v], visited):
match[v] = u
return True
return False
result = 0
for u in range(n):
visited = [False] * m
if dfs(u, visited):
result += 1
return result
2. Bipartite Graph Validation with Custom Logic
def validate_bipartite_assignment(assignments, conflicts):
"""
Validate if assignment is bipartite given conflict pairs
assignments: list of items to assign
conflicts: list of (item1, item2) that cannot be in same group
"""
from collections import defaultdict
graph = defaultdict(list)
for u, v in conflicts:
graph[u].append(v)
graph[v].append(u)
colors = {}
def can_color(item, color):
if item in colors:
return colors[item] == color
colors[item] = color
for conflict_item in graph[item]:
if not can_color(conflict_item, 1 - color):
return False
return True
for item in assignments:
if item not in colors:
if not can_color(item, 0):
return False, {}
# Return partition
group_a = [item for item, color in colors.items() if color == 0]
group_b = [item for item, color in colors.items() if color == 1]
return True, {"Group A": group_a, "Group B": group_b}
Performance Comparison
| Approach | Time | Space | Best Use Case |
|---|---|---|---|
| BFS | O(V+E) | O(V) | Level-by-level processing |
| DFS | O(V+E) | O(V) | Simple recursive solution |
| Union-Find | O(E⋅α(V)) | O(V) | Dynamic conflict detection |
| Grid-specific | O(R⋅C) | O(R⋅C) | 2D grid problems |
Common Patterns & Tips
Pattern Recognition:
- Graph coloring → Bipartite check
- Conflict/compatibility → Build conflict graph
- Two groups assignment → Bipartite partition
- Matching problems → Bipartite matching
Implementation Tips:
- Always handle disconnected components
- Use 0/1 or -1/1 for colors consistently
- Check conflicts immediately when coloring
- Consider Union-Find for dynamic scenarios
Edge Cases:
- Empty graph (bipartite by definition)
- Single node (bipartite)
- No edges (bipartite)
- Self-loops (not bipartite if exists)
- Disconnected components (check all)
Template 7: Shortest Path Algorithms
Overview: Shortest Path Algorithm Comparison
| Algorithm | Time | Space | Edge Weights | Use Case | Best For |
|---|---|---|---|---|---|
| BFS | O(V+E) | O(V) | Unweighted | Single source | Unweighted graphs |
| Dijkstra | O((V+E)logV) | O(V) | Non-negative | Single source | Non-negative weights |
| Bellman-Ford | O(V⋅E) | O(V) | Any (including negative) | Single source | Negative edges, detect cycles |
| Floyd-Warshall | O(V³) | O(V²) | Any (including negative) | All pairs | Dense graphs, all-pairs |
| SPFA | O(V⋅E) avg | O(V) | Any | Single source | Sparse graphs with negative |
When to Use Each:
- BFS: Unweighted graphs, shortest path in terms of number of edges
- Dijkstra: Non-negative weights, need optimal performance
- Bellman-Ford: Negative weights exist, need to detect negative cycles
- Floyd-Warshall: Need all-pairs shortest paths, small dense graph
- SPFA (Shortest Path Faster Algorithm): Bellman-Ford optimization, average O(E)
7.1) Bellman-Ford Algorithm
Core Concept: Bellman-Ford finds shortest paths from a single source to all vertices, even with negative edge weights. It can also detect negative cycles.
Key Insight:
- Relaxes all edges V-1 times (V = number of vertices)
- After V-1 iterations, shortest paths are found (if no negative cycle)
- One more iteration detects negative cycles
Algorithm Steps:
- Initialize distances:
dist[source] = 0, all others = ∞ - Relax all edges V-1 times:
- For each edge (u, v, weight):
- If
dist[u] + weight < dist[v]: updatedist[v]
- If
- For each edge (u, v, weight):
- Check for negative cycles:
- If any edge can still be relaxed, negative cycle exists
Why V-1 Iterations?
In a graph with V vertices, the shortest path between any two vertices
contains at most V-1 edges (no cycles).
After k iterations, we have the shortest path using at most k edges.
After V-1 iterations, we have the shortest paths for all pairs.
Python Implementation
# Bellman-Ford Algorithm
def bellman_ford(n, edges, source):
"""
Find shortest paths from source to all vertices.
Can handle negative weights and detect negative cycles.
Args:
n: number of vertices (0 to n-1)
edges: list of (u, v, weight) tuples
source: starting vertex
Returns:
dist: array of shortest distances (or None if negative cycle)
Time: O(V⋅E)
Space: O(V)
"""
# Initialize distances
dist = [float('inf')] * n
dist[source] = 0
# Relax all edges V-1 times
for _ in range(n - 1):
updated = False
for u, v, weight in edges:
if dist[u] != float('inf') and dist[u] + weight < dist[v]:
dist[v] = dist[u] + weight
updated = True
# Early termination if no updates
if not updated:
break
# Check for negative cycles
for u, v, weight in edges:
if dist[u] != float('inf') and dist[u] + weight < dist[v]:
return None # Negative cycle detected
return dist
# Example usage:
# edges = [(0, 1, 4), (0, 2, 5), (1, 2, -3), (2, 3, 4)]
# result = bellman_ford(4, edges, 0)
# Output: [0, 4, 1, 5] (shortest distances from vertex 0)
Java Implementation
// Bellman-Ford Algorithm
/**
* LC 787 - Cheapest Flights Within K Stops (Bellman-Ford variant)
*
* time = O(V⋅E) or O(K⋅E) for K iterations
* space = O(V)
*/
class Solution {
public int[] bellmanFord(int n, int[][] edges, int src) {
// Initialize distances
int[] dist = new int[n];
Arrays.fill(dist, Integer.MAX_VALUE);
dist[src] = 0;
// Relax edges V-1 times
for (int i = 0; i < n - 1; i++) {
boolean updated = false;
for (int[] edge : edges) {
int u = edge[0];
int v = edge[1];
int weight = edge[2];
if (dist[u] != Integer.MAX_VALUE && dist[u] + weight < dist[v]) {
dist[v] = dist[u] + weight;
updated = true;
}
}
// Early termination
if (!updated) break;
}
// Check for negative cycle
for (int[] edge : edges) {
int u = edge[0];
int v = edge[1];
int weight = edge[2];
if (dist[u] != Integer.MAX_VALUE && dist[u] + weight < dist[v]) {
return null; // Negative cycle detected
}
}
return dist;
}
}
Visual Example
Graph with negative edge:
0 --4--> 1
| / |
5 -3 |
| / 4
v v v
2 --------> 3
Edges: [(0,1,4), (0,2,5), (1,2,-3), (2,3,4)]
Iteration 0 (Initial):
dist = [0, ∞, ∞, ∞]
Iteration 1:
Edge (0,1,4): dist[1] = 0 + 4 = 4
Edge (0,2,5): dist[2] = 0 + 5 = 5
Edge (1,2,-3): dist[2] = min(5, 4-3) = 1
Edge (2,3,4): dist[3] = 1 + 4 = 5
dist = [0, 4, 1, 5]
Iteration 2:
Edge (1,2,-3): dist[2] = min(1, 4-3) = 1 (no change)
Edge (2,3,4): dist[3] = min(5, 1+4) = 5 (no change)
No updates → Early termination
Final: dist = [0, 4, 1, 5] ✓
Negative Cycle Detection
# Example with negative cycle
def detect_negative_cycle(n, edges):
"""
Detect if graph contains a negative cycle.
Time: O(V⋅E)
Space: O(V)
"""
# Pick arbitrary source (negative cycle affects all paths)
source = 0
dist = [float('inf')] * n
dist[source] = 0
# Relax V-1 times
for _ in range(n - 1):
for u, v, weight in edges:
if dist[u] != float('inf') and dist[u] + weight < dist[v]:
dist[v] = dist[u] + weight
# Check if can still relax
for u, v, weight in edges:
if dist[u] != float('inf') and dist[u] + weight < dist[v]:
return True # Negative cycle exists
return False
# Example: edges = [(0,1,1), (1,2,-3), (2,0,1)]
# Forms cycle: 0→1→2→0 with total weight 1-3+1 = -1
# Returns: True
7.2) Floyd-Warshall Algorithm
Core Concept: Floyd-Warshall finds all-pairs shortest paths in a weighted graph using dynamic programming. Works with negative weights but not negative cycles.
Key Insight:
- DP approach: For each pair (i, j), try all intermediate vertices k
dist[i][j] = min(dist[i][j], dist[i][k] + dist[k][j])- After considering all k, we have shortest paths for all pairs
Algorithm Steps:
- Initialize
dist[i][j]:dist[i][i] = 0(same vertex)dist[i][j] = weight(i, j)if edge existsdist[i][j] = ∞otherwise
- For each intermediate vertex k (0 to n-1):
- For each pair (i, j):
dist[i][j] = min(dist[i][j], dist[i][k] + dist[k][j])
- For each pair (i, j):
Why It Works:
After considering vertices {0, 1, ..., k} as intermediate:
dist[i][j] = shortest path from i to j using only vertices {0...k}
When k = n-1, we've considered all possible intermediates
→ dist[i][j] = shortest path from i to j
Python Implementation
# Floyd-Warshall Algorithm
def floyd_warshall(n, edges):
"""
Find all-pairs shortest paths.
Args:
n: number of vertices (0 to n-1)
edges: list of (u, v, weight) tuples
Returns:
dist: 2D array where dist[i][j] = shortest path from i to j
Time: O(V³)
Space: O(V²)
"""
# Initialize distance matrix
dist = [[float('inf')] * n for _ in range(n)]
# Distance from vertex to itself is 0
for i in range(n):
dist[i][i] = 0
# Fill in edge weights
for u, v, weight in edges:
dist[u][v] = weight
# For undirected graphs, uncomment:
# dist[v][u] = weight
# Dynamic programming: try all intermediate vertices
for k in range(n):
for i in range(n):
for j in range(n):
# Can we improve path i→j by going through k?
if dist[i][k] != float('inf') and dist[k][j] != float('inf'):
dist[i][j] = min(dist[i][j], dist[i][k] + dist[k][j])
# Check for negative cycles
for i in range(n):
if dist[i][i] < 0:
return None # Negative cycle detected
return dist
# Example usage:
# edges = [(0,1,3), (1,2,1), (0,2,7), (2,3,2)]
# result = floyd_warshall(4, edges)
# result[i][j] = shortest distance from i to j
Java Implementation
// Floyd-Warshall Algorithm
/**
* LC 1334 - Find the City With the Smallest Number of Neighbors
*
* time = O(V³)
* space = O(V²)
*/
class Solution {
public int[][] floydWarshall(int n, int[][] edges) {
// Initialize distance matrix
int[][] dist = new int[n][n];
// Fill with infinity
for (int i = 0; i < n; i++) {
Arrays.fill(dist[i], Integer.MAX_VALUE / 2); // Avoid overflow
dist[i][i] = 0; // Distance to self is 0
}
// Add edge weights
for (int[] edge : edges) {
int u = edge[0];
int v = edge[1];
int weight = edge[2];
dist[u][v] = weight;
dist[v][u] = weight; // For undirected graph
}
// Floyd-Warshall: try all intermediate vertices
for (int k = 0; k < n; k++) {
for (int i = 0; i < n; i++) {
for (int j = 0; j < n; j++) {
// Relax through vertex k
dist[i][j] = Math.min(dist[i][j], dist[i][k] + dist[k][j]);
}
}
}
return dist;
}
}
Visual Example
Graph:
0 --3--> 1
| |
7 1
| |
v v
2 <--2-- 3
Initial distance matrix:
0 1 2 3
0 [ 0 3 7 ∞ ]
1 [ ∞ 0 ∞ 1 ]
2 [ ∞ ∞ 0 ∞ ]
3 [ ∞ ∞ 2 0 ]
After k=0 (intermediate vertex 0):
No changes (0 is only source, not intermediate)
After k=1 (intermediate vertex 1):
dist[0][3] = min(∞, dist[0][1] + dist[1][3]) = min(∞, 3+1) = 4
0 1 2 3
0 [ 0 3 7 4 ]
1 [ ∞ 0 ∞ 1 ]
2 [ ∞ ∞ 0 ∞ ]
3 [ ∞ ∞ 2 0 ]
After k=2:
No improvements
After k=3 (intermediate vertex 3):
dist[0][2] = min(7, dist[0][3] + dist[3][2]) = min(7, 4+2) = 6
dist[1][2] = min(∞, dist[1][3] + dist[3][2]) = min(∞, 1+2) = 3
Final:
0 1 2 3
0 [ 0 3 6 4 ]
1 [ ∞ 0 3 1 ]
2 [ ∞ ∞ 0 ∞ ]
3 [ ∞ ∞ 2 0 ]
Path Reconstruction
# Floyd-Warshall with path reconstruction
def floyd_warshall_with_path(n, edges):
"""
Find all-pairs shortest paths and reconstruct paths.
Returns:
dist: shortest distances
next_vertex: for path reconstruction
"""
dist = [[float('inf')] * n for _ in range(n)]
next_vertex = [[None] * n for _ in range(n)]
# Initialize
for i in range(n):
dist[i][i] = 0
for u, v, weight in edges:
dist[u][v] = weight
next_vertex[u][v] = v
# Floyd-Warshall
for k in range(n):
for i in range(n):
for j in range(n):
if dist[i][k] + dist[k][j] < dist[i][j]:
dist[i][j] = dist[i][k] + dist[k][j]
next_vertex[i][j] = next_vertex[i][k]
return dist, next_vertex
def reconstruct_path(next_vertex, start, end):
"""Reconstruct shortest path from start to end."""
if next_vertex[start][end] is None:
return []
path = [start]
current = start
while current != end:
current = next_vertex[current][end]
path.append(current)
return path
# Example:
# dist, next_v = floyd_warshall_with_path(4, edges)
# path = reconstruct_path(next_v, 0, 3)
# Output: [0, 1, 3] (path from 0 to 3)
7.3) Classic LeetCode Problems
| Problem | LC# | Algorithm | Difficulty | Key Insight |
|---|---|---|---|---|
| Network Delay Time | 743 | Bellman-Ford / Dijkstra | Medium | Single source shortest path |
| Cheapest Flights K Stops | 787 | Bellman-Ford (K iterations) | Medium | Limit iterations to K+1 |
| Find the City | 1334 | Floyd-Warshall | Medium | All-pairs, count neighbors |
| Course Schedule III | 630 | Bellman-Ford variant | Hard | With constraints |
| Minimum Cost to Reach Destination | 1928 | Bellman-Ford / Dijkstra | Hard | Modified edge costs |
| Bellman-Ford | 1724 | Bellman-Ford | Medium | Check negative cycles |
7.4) Performance Comparison & When to Use
Bellman-Ford vs Dijkstra:
| Aspect | Bellman-Ford | Dijkstra |
|---|---|---|
| Time | O(V⋅E) slower | O((V+E)logV) faster |
| Edge Weights | Any (including negative) | Non-negative only |
| Negative Cycles | Can detect | Cannot handle |
| Implementation | Simpler | More complex (priority queue) |
| Use Case | Negative weights, cycle detection | Optimal for non-negative |
Floyd-Warshall vs Running Dijkstra V times:
| Aspect | Floyd-Warshall | V × Dijkstra |
|---|---|---|
| Time | O(V³) | O(V²⋅E⋅logV) |
| Space | O(V²) | O(V) |
| Code Complexity | Simple (3 loops) | More complex |
| Best For | Dense graphs, small V | Sparse graphs, large V |
Decision Tree:
Need shortest paths?
├─ Single source
│ ├─ Negative weights? → Bellman-Ford
│ └─ Non-negative? → Dijkstra (faster)
│
└─ All pairs
├─ Dense graph or small V? → Floyd-Warshall
└─ Sparse graph or large V? → Run Dijkstra V times
7.5) Interview Tips
1. Algorithm Selection:
Q: "Find shortest path from A to B"
→ Clarify: Negative weights? → Bellman-Ford : Dijkstra
Q: "Find shortest paths between all pairs"
→ Ask: Graph density? → Dense: Floyd-Warshall, Sparse: Dijkstra×V
Q: "Detect if negative cycle exists"
→ Use: Bellman-Ford (only algorithm that detects this)
2. Common Mistakes:
- Bellman-Ford: Forgetting to check
dist[u] != INFbefore relaxation - Floyd-Warshall: Wrong loop order (must be k→i→j, not i→j→k)
- Both: Not handling unreachable vertices (dist = INF)
- Negative cycles: Not checking after main algorithm
3. Optimization Tips:
# Bellman-Ford: Early termination
for iteration in range(n - 1):
updated = False
for edge in edges:
if relax(edge):
updated = True
if not updated:
break # No more improvements possible
# Floyd-Warshall: Only update if improves
if dist[i][k] + dist[k][j] < dist[i][j]:
dist[i][j] = dist[i][k] + dist[k][j]
4. Edge Cases:
- Graph with no edges (all distances = INF except self)
- Negative cycle (Bellman-Ford returns None)
- Disconnected graph (some distances remain INF)
- Self-loops with negative weight (negative cycle)
5. Talking Points:
- “Bellman-Ford trades time for flexibility with negative weights”
- “Floyd-Warshall is DP: optimal substructure through intermediate vertices”
- “V-1 iterations because longest simple path has V-1 edges”
- “Floyd-Warshall is simple but O(V³) limits scalability”
Template 8: Tarjan’s Algorithm (Graph Connectivity)
Overview: Tarjan’s algorithm is a DFS-based technique for finding critical graph structures:
- Strongly Connected Components (SCC) - Maximal sets of mutually reachable vertices (directed graphs)
- Bridges - Edges whose removal disconnects the graph (undirected graphs)
- Articulation Points (Cut Vertices) - Vertices whose removal disconnects the graph (undirected graphs)
Core Concept: Uses DFS with two key arrays:
disc[v]: Discovery time of vertex v (when first visited)low[v]: Lowest discovery time reachable from v’s subtree
Time Complexity: O(V + E) - single DFS traversal Space Complexity: O(V) - recursion stack + arrays
8.1) Strongly Connected Components (SCC)
Definition: In a directed graph, an SCC is a maximal set of vertices where every vertex is reachable from every other vertex in the set.
Key Insight:
- Use a stack to track vertices in current DFS path
- When
low[v] == disc[v], v is the root of an SCC - Pop all vertices from stack until v to get complete SCC
Algorithm Steps:
- Initialize
disc[],low[], and stack - DFS from each unvisited vertex
- For each vertex v:
- Set
disc[v] = low[v] = timer++ - Push v onto stack
- For each neighbor u:
- If unvisited: DFS(u), update
low[v] = min(low[v], low[u]) - If u on stack: update
low[v] = min(low[v], disc[u])
- If unvisited: DFS(u), update
- If
low[v] == disc[v]: pop stack until v to form SCC
- Set
Python Implementation
# Tarjan's Algorithm for SCC
def tarjan_scc(n, graph):
"""
Find all strongly connected components using Tarjan's algorithm.
Args:
n: number of vertices (0 to n-1)
graph: adjacency list (directed graph)
Returns:
List of SCCs, where each SCC is a list of vertices
Time: O(V + E)
Space: O(V)
"""
disc = [-1] * n # Discovery times
low = [-1] * n # Lowest reachable
on_stack = [False] * n
stack = []
sccs = []
timer = [0] # Use list for mutability
def dfs(v):
# Initialize discovery time and low value
disc[v] = low[v] = timer[0]
timer[0] += 1
stack.append(v)
on_stack[v] = True
# Explore neighbors
for u in graph[v]:
if disc[u] == -1:
# Unvisited neighbor
dfs(u)
low[v] = min(low[v], low[u])
elif on_stack[u]:
# Back edge to vertex on stack
low[v] = min(low[v], disc[u])
# If v is a root of SCC, pop the SCC
if low[v] == disc[v]:
scc = []
while True:
u = stack.pop()
on_stack[u] = False
scc.append(u)
if u == v:
break
sccs.append(scc)
# Run DFS from each unvisited vertex
for i in range(n):
if disc[i] == -1:
dfs(i)
return sccs
# Example:
# graph = {0: [1], 1: [2], 2: [0, 3], 3: [4], 4: [5], 5: [3]}
# 0 → 1 → 2
# ↑ ↓
# └───────┘ 3 ⇄ 4 → 5
# ↑ ↓
# └───┘
# SCCs: [[0, 2, 1], [3, 5, 4]]
Java Implementation
// Tarjan's SCC Algorithm
/**
* LC 1192 - Critical Connections in a Network (related)
*
* time = O(V + E)
* space = O(V)
*/
class TarjanSCC {
private int timer = 0;
private int[] disc;
private int[] low;
private boolean[] onStack;
private Stack<Integer> stack;
private List<List<Integer>> sccs;
public List<List<Integer>> findSCCs(int n, List<List<Integer>> graph) {
disc = new int[n];
low = new int[n];
onStack = new boolean[n];
stack = new Stack<>();
sccs = new ArrayList<>();
Arrays.fill(disc, -1);
Arrays.fill(low, -1);
// DFS from each unvisited vertex
for (int i = 0; i < n; i++) {
if (disc[i] == -1) {
dfs(i, graph);
}
}
return sccs;
}
private void dfs(int v, List<List<Integer>> graph) {
// Initialize
disc[v] = low[v] = timer++;
stack.push(v);
onStack[v] = true;
// Explore neighbors
for (int u : graph.get(v)) {
if (disc[u] == -1) {
// Unvisited
dfs(u, graph);
low[v] = Math.min(low[v], low[u]);
} else if (onStack[u]) {
// Back edge
low[v] = Math.min(low[v], disc[u]);
}
}
// Root of SCC found
if (low[v] == disc[v]) {
List<Integer> scc = new ArrayList<>();
while (true) {
int u = stack.pop();
onStack[u] = false;
scc.add(u);
if (u == v) break;
}
sccs.add(scc);
}
}
}
8.2) Finding Bridges (Critical Connections)
Definition: A bridge is an edge whose removal increases the number of connected components (disconnects the graph).
Key Insight:
- Edge (u, v) is a bridge if
low[v] > disc[u] - Means v cannot reach any vertex discovered before u without using edge (u, v)
Algorithm Steps:
- Run DFS with
disc[]andlow[] - For each edge (u, v) in DFS tree:
- If
low[v] > disc[u]: (u, v) is a bridge
- If
Python Implementation
# Tarjan's Algorithm for Bridges
def find_bridges(n, edges):
"""
Find all bridges (critical connections) in an undirected graph.
Args:
n: number of vertices
edges: list of [u, v] edges
Returns:
List of bridges (critical edges)
Time: O(V + E)
Space: O(V + E)
"""
# Build adjacency list
graph = [[] for _ in range(n)]
for u, v in edges:
graph[u].append(v)
graph[v].append(u)
disc = [-1] * n
low = [-1] * n
bridges = []
timer = [0]
def dfs(v, parent):
disc[v] = low[v] = timer[0]
timer[0] += 1
for u in graph[v]:
if u == parent:
# Skip edge to parent (undirected graph)
continue
if disc[u] == -1:
# Unvisited neighbor
dfs(u, v)
low[v] = min(low[v], low[u])
# Check if (v, u) is a bridge
if low[u] > disc[v]:
bridges.append([v, u])
else:
# Back edge
low[v] = min(low[v], disc[u])
# Run DFS from each component
for i in range(n):
if disc[i] == -1:
dfs(i, -1)
return bridges
# Example:
# n = 4, edges = [[0,1],[1,2],[2,0],[1,3]]
#
# 0 --- 1 --- 3
# \ /
# \ /
# 2
#
# Bridge: [1, 3] (removing this disconnects 3 from rest)
Java Implementation
// LC 1192 - Critical Connections in a Network
/**
* time = O(V + E)
* space = O(V + E)
*/
class Solution {
private int timer = 0;
private int[] disc;
private int[] low;
private List<List<Integer>> bridges;
public List<List<Integer>> criticalConnections(int n, List<List<Integer>> connections) {
// Build adjacency list
List<List<Integer>> graph = new ArrayList<>();
for (int i = 0; i < n; i++) {
graph.add(new ArrayList<>());
}
for (List<Integer> conn : connections) {
int u = conn.get(0);
int v = conn.get(1);
graph.get(u).add(v);
graph.get(v).add(u);
}
disc = new int[n];
low = new int[n];
bridges = new ArrayList<>();
Arrays.fill(disc, -1);
// DFS from vertex 0 (graph is connected in this problem)
dfs(0, -1, graph);
return bridges;
}
private void dfs(int v, int parent, List<List<Integer>> graph) {
disc[v] = low[v] = timer++;
for (int u : graph.get(v)) {
if (u == parent) continue; // Skip parent edge
if (disc[u] == -1) {
// Unvisited
dfs(u, v, graph);
low[v] = Math.min(low[v], low[u]);
// Check for bridge
if (low[u] > disc[v]) {
bridges.add(Arrays.asList(v, u));
}
} else {
// Back edge
low[v] = Math.min(low[v], disc[u]);
}
}
}
}
8.3) Finding Articulation Points (Cut Vertices)
Definition: An articulation point is a vertex whose removal increases the number of connected components.
Key Insight:
- Vertex u is an articulation point if:
- Root of DFS tree: has 2+ children
- Non-root: has a child v where
low[v] >= disc[u]
Algorithm Steps:
- Run DFS with
disc[]andlow[] - For each vertex u:
- If root: count children, articulation point if ≥ 2
- If non-root: check if any child v has
low[v] >= disc[u]
Python Implementation
# Tarjan's Algorithm for Articulation Points
def find_articulation_points(n, edges):
"""
Find all articulation points (cut vertices).
Args:
n: number of vertices
edges: list of [u, v] edges
Returns:
Set of articulation points
Time: O(V + E)
Space: O(V + E)
"""
# Build adjacency list
graph = [[] for _ in range(n)]
for u, v in edges:
graph[u].append(v)
graph[v].append(u)
disc = [-1] * n
low = [-1] * n
ap = set() # Articulation points
timer = [0]
def dfs(v, parent):
children = 0
disc[v] = low[v] = timer[0]
timer[0] += 1
for u in graph[v]:
if u == parent:
continue
if disc[u] == -1:
# Unvisited child
children += 1
dfs(u, v)
low[v] = min(low[v], low[u])
# Check if v is articulation point
# Case 1: Root with 2+ children
if parent == -1 and children > 1:
ap.add(v)
# Case 2: Non-root with child that can't reach ancestor
if parent != -1 and low[u] >= disc[v]:
ap.add(v)
else:
# Back edge
low[v] = min(low[v], disc[u])
# Run DFS from each component
for i in range(n):
if disc[i] == -1:
dfs(i, -1)
return list(ap)
# Example:
# n = 5, edges = [[0,1],[1,2],[2,0],[1,3],[3,4]]
#
# 0 --- 1 --- 3 --- 4
# \ /
# \ /
# 2
#
# Articulation points: [1, 3]
# (Removing 1 disconnects {0,2} from {3,4})
# (Removing 3 disconnects 4 from rest)
Java Implementation
// Articulation Points Algorithm
/**
* time = O(V + E)
* space = O(V + E)
*/
class ArticulationPoints {
private int timer = 0;
private int[] disc;
private int[] low;
private Set<Integer> ap;
public List<Integer> findArticulationPoints(int n, int[][] edges) {
// Build adjacency list
List<List<Integer>> graph = new ArrayList<>();
for (int i = 0; i < n; i++) {
graph.add(new ArrayList<>());
}
for (int[] edge : edges) {
int u = edge[0];
int v = edge[1];
graph.get(u).add(v);
graph.get(v).add(u);
}
disc = new int[n];
low = new int[n];
ap = new HashSet<>();
Arrays.fill(disc, -1);
// DFS from each component
for (int i = 0; i < n; i++) {
if (disc[i] == -1) {
dfs(i, -1, graph);
}
}
return new ArrayList<>(ap);
}
private void dfs(int v, int parent, List<List<Integer>> graph) {
int children = 0;
disc[v] = low[v] = timer++;
for (int u : graph.get(v)) {
if (u == parent) continue;
if (disc[u] == -1) {
children++;
dfs(u, v, graph);
low[v] = Math.min(low[v], low[u]);
// Root with 2+ children
if (parent == -1 && children > 1) {
ap.add(v);
}
// Non-root with blocking child
if (parent != -1 && low[u] >= disc[v]) {
ap.add(v);
}
} else {
// Back edge
low[v] = Math.min(low[v], disc[u]);
}
}
}
}
8.4) Visual Example: Tarjan’s Algorithm Walkthrough
Graph (Directed):
0 → 1 → 2
↑ ↓
└───────┘ 3 ⇄ 4
↓ ↑
5 ──┘
DFS Traversal:
Step 1: Start at 0
disc[0] = low[0] = 0
stack = [0]
Step 2: Visit 1 from 0
disc[1] = low[1] = 1
stack = [0, 1]
Step 3: Visit 2 from 1
disc[2] = low[2] = 2
stack = [0, 1, 2]
Step 4: Back edge 2→0 (0 already on stack)
low[2] = min(2, disc[0]) = 0
Backtrack to 1: low[1] = min(1, low[2]) = 0
Backtrack to 0: low[0] = min(0, low[1]) = 0
Step 5: At 0, low[0] == disc[0] → SCC found!
Pop stack: [2, 1, 0]
SCC #1: {0, 1, 2}
Step 6: Start at 3
disc[3] = low[3] = 3
stack = [3]
Step 7: Visit 4 from 3
disc[4] = low[4] = 4
stack = [3, 4]
Step 8: Visit 5 from 4
disc[5] = low[5] = 5
stack = [3, 4, 5]
Step 9: Edge 5→3 (back edge)
low[5] = min(5, disc[3]) = 3
Backtrack to 4: low[4] = min(4, low[5]) = 3
Backtrack to 3: low[3] = min(3, low[4]) = 3
Step 10: At 3, low[3] == disc[3] → SCC found!
Pop stack: [5, 4, 3]
SCC #2: {3, 4, 5}
Final SCCs: [{0,1,2}, {3,4,5}]
8.5) Classic LeetCode Problems
| Problem | LC# | Variant | Difficulty | Key Insight |
|---|---|---|---|---|
| Critical Connections in Network | 1192 | Bridges | Hard | Find all bridges using Tarjan |
| Number of Provinces | 547 | Basic connectivity | Medium | Count connected components |
| Redundant Connection | 684 | Cycle detection | Medium | Find edge creating cycle |
| Redundant Connection II | 685 | Directed graph | Hard | SCC + cycle in directed graph |
| Minimum Number of Vertices | 1557 | SCC sources | Medium | Find vertices with no incoming |
8.6) Comparison: Tarjan vs Kosaraju for SCC
| Aspect | Tarjan’s Algorithm | Kosaraju’s Algorithm |
|---|---|---|
| Passes | Single DFS | Two DFS passes |
| Time | O(V + E) | O(V + E) |
| Space | O(V) stack | O(V) + transpose graph |
| Complexity | More complex (one pass) | Simpler (two passes) |
| Extra Space | Stack for SCC | Reversed graph |
| Preference | More efficient (one pass) | Easier to understand |
8.7) Interview Tips
1. Recognition Patterns:
"critical connections" → Bridges (Tarjan)
"strongly connected" → SCC (Tarjan or Kosaraju)
"cut vertices" → Articulation points (Tarjan)
"remove vertex/edge disconnects graph" → Articulation/Bridge
2. Key Differences:
SCC: Directed graph, maximal mutually reachable sets
Bridges: Undirected graph, critical edges
Articulation Points: Undirected graph, critical vertices
low[v] == disc[v] → Root of SCC (directed)
low[v] > disc[u] → (u,v) is bridge (undirected)
low[v] >= disc[u] → u is articulation point (undirected)
3. Common Mistakes:
- Forgetting to skip parent edge in undirected graphs
- Wrong condition for articulation point (root vs non-root)
- Not using
on_stackarray for SCC (leads to incorrect SCCs) - Confusing
disc[u]vslow[u]in back edge updates
4. Template to Memorize:
def tarjan_template(v, parent=-1):
disc[v] = low[v] = timer
timer += 1
for u in graph[v]:
if u == parent: # Undirected graphs only
continue
if disc[u] == -1:
# Tree edge
dfs(u, v)
low[v] = min(low[v], low[u])
# Check condition here (bridge, AP, etc.)
else:
# Back edge
low[v] = min(low[v], disc[u]) # Or check on_stack for SCC
5. Talking Points:
- “Tarjan’s uses single DFS with discovery times”
- “low[v] tracks earliest reachable vertex from v’s subtree”
- “Bridges/APs indicate critical graph structure”
- “SCCs represent maximal strongly connected regions”
Problems by Pattern
Graph Traversal Problems
| Problem | LC # | Key Technique | Difficulty |
|---|---|---|---|
| Number of Islands | 200 | DFS/BFS on grid | Medium |
| Max Area of Island | 695 | DFS with counting | Medium |
| Clone Graph | 133 | BFS/DFS with map | Medium |
| Pacific Atlantic Water | 417 | Multi-source DFS | Medium |
| Word Ladder | 127 | BFS shortest path | Hard |
| Surrounded Regions | 130 | DFS from boundary | Medium |
Shortest Path Problems
| Problem | LC # | Key Technique | Difficulty |
|---|---|---|---|
| Network Delay Time | 743 | Dijkstra | Medium |
| Cheapest Flights K Stops | 787 | Modified Dijkstra | Medium |
| Path with Min Effort | 1631 | Dijkstra on grid | Medium |
| Bus Routes | 815 | BFS on routes | Hard |
| Shortest Path Binary Matrix | 1091 | BFS | Medium |
Union-Find Problems
| Problem | LC # | Key Technique | Difficulty |
|---|---|---|---|
| Number of Connected Components | 323 | Basic Union-Find | Medium |
| Redundant Connection | 684 | Detect cycle | Medium |
| Accounts Merge | 721 | Union-Find with map | Medium |
| Number of Provinces | 547 | Union-Find or DFS | Medium |
| Satisfiability of Equality | 990 | Union-Find | Medium |
Topological Sort Problems
| Problem | LC # | Key Technique | Difficulty |
|---|---|---|---|
| Course Schedule | 207 | Cycle detection | Medium |
| Course Schedule II | 210 | Topological order | Medium |
| Alien Dictionary | 269 | Build graph + sort | Hard |
| Minimum Height Trees | 310 | Leaf removal | Medium |
| Parallel Courses | 1136 | Level-wise BFS | Medium |
Bipartite Problems
| Problem | LC # | Key Technique | Difficulty |
|---|---|---|---|
| Is Graph Bipartite | 785 | BFS coloring | Medium |
| Possible Bipartition | 886 | DFS coloring | Medium |
Advanced Graph Problems
| Problem | LC # | Key Technique | Difficulty |
|---|---|---|---|
| Critical Connections | 1192 | Tarjan’s algorithm | Hard |
| Find Eventual Safe States | 802 | Cycle detection | Medium |
| Reconstruct Itinerary | 332 | Hierholzer’s algorithm | Hard |
| Minimum Spanning Tree | 1135 | Kruskal/Prim | Medium |
1-1-1) Number of Islands
- LC 200
// java
void dfs(char[][] grid, int r, int c){
int nr = grid.length;
int nc = grid[0].length;
if (r < 0 || c < 0 || r >= nr || c >= nc || grid[r][c] == '0') {
return;
}
grid[r][c] = '0';
/** NOTE here !!!*/
dfs_1(grid, r - 1, c);
dfs_1(grid, r + 1, c);
dfs_1(grid, r, c - 1);
dfs_1(grid, r, c + 1);
}
public int numIslands_1(char[][] grid) {
if (grid == null || grid.length == 0) {
return 0;
}
int nr = grid.length;
int nc = grid[0].length;
int num_islands = 0;
for (int r = 0; r < nr; ++r) {
for (int c = 0; c < nc; ++c) {
if (grid[r][c] == '1') {
++num_islands;
dfs_1(grid, r, c);
}
}
}
return num_islands;
}
1-1-2) Max Area of Island
- LC 695
// java
int[][] grid;
boolean[][] seen;
public int area(int r, int c) {
if (r < 0 || r >= grid.length || c < 0 || c >= grid[0].length ||
seen[r][c] || grid[r][c] == 0)
return 0;
seen[r][c] = true;
/** NOTE !!!*/
return (1 + area(r+1, c) + area(r-1, c)
+ area(r, c-1) + area(r, c+1));
}
public int maxAreaOfIsland_1(int[][] grid) {
this.grid = grid;
seen = new boolean[grid.length][grid[0].length];
int ans = 0;
for (int r = 0; r < grid.length; r++) {
for (int c = 0; c < grid[0].length; c++) {
ans = Math.max(ans, area(r, c));
}
}
return ans;
}
2) LC Example
2-1) Closest Leaf in a Binary Tree
# 742 Closest Leaf in a Binary Tree
import collections
class Solution:
# search via DFS
def findClosestLeaf(self, root, k):
self.start = None
self.buildGraph(root, None, k)
q, visited = [root], set()
#q, visited = [self.start], set() # need to validate this
self.graph = collections.defaultdict(list)
while q:
for i in range(len(q)):
cur = q.pop(0) # this is dfs
# add cur to visited, NOT to visit this node again
visited.add(cur)
### NOTICE HERE
# if not cur.left and not cur.right: means this is the leaf (HAS NO ANY left/right node) of the tree
# so the first value of this is what we want, just return cur.val as answer directly
if not cur.left and not cur.right:
# return the answer
return cur.val
# if not find the leaf, then go through all neighbors of current node, and search again
for node in self.graph:
if node not in visited: # need to check if "if node not in visited" or "if node in visited"
q.append(node)
# build graph via DFS
# node : current node
# parent : parent of current node
def buildGraph(self, node, parent, k):
if not node:
return
# if node.val == k, THEN GET THE start point FROM current "node",
# then build graph based on above
if node.val == k:
self.start = node
if parent:
self.graph[node].append(parent)
self.graph[parent].append(node)
self.buildGraph(node.left, node, k)
self.buildGraph(node.right, node, k)
2-2) Number of Connected Components in an Undirected Graph
# LC 323 Number of Connected Components in an Undirected Graph
# V0
# IDEA : DFS
class Solution:
def countComponents(self, n, edges):
def helper(u):
if u in pair:
for v in pair[u]:
if v not in visited:
visited.add(v)
helper(v)
pair = collections.defaultdict(set)
for u,v in edges:
pair[u].add(v)
pair[v].add(u)
count = 0
visited = set()
for i in range(n):
if i not in visited:
helper(i)
count+=1
return count
2-3) Clone Graph
# LC 133. Clone Graph
# V0
# IDEA : BFS
class Solution(object):
def cloneGraph(self, node):
if not node:
return
q = [node]
"""
NOTE !!! : we init res as Node(node.val, [])
-> since Node has structure as below :
class Node:
def __init__(self, val = 0, neighbors = None):
self.val = val
self.neighbors = neighbors if neighbors is not None else []
"""
res = Node(node.val, [])
"""
NOTE !!! : we use dict as visited,
and we use node as visited dict key
"""
visited = dict()
visited[node] = res
while q:
#t = q.pop(0) # this works as well
t = q.pop(-1)
if not t:
continue
for n in t.neighbors:
if n not in visited:
"""
NOTE !!! : we need to
-> use n as visited key
-> use Node(n.val, []) as visited value
"""
visited[n] = Node(n.val, [])
q.append(n)
"""
NOTE !!!
-> we need to append visited[n] to visited[t].neighbors
"""
visited[t].neighbors.append(visited[n])
return res
# V0
# IDEA : DFS
# NOTE :
# -> 1) we init node via : node_copy = Node(node.val, [])
# -> 2) we copy graph via dict
class Solution(object):
def cloneGraph(self, node):
"""
:type node: Node
:rtype: Node
"""
node_copy = self.dfs(node, dict())
return node_copy
def dfs(self, node, hashd):
if not node: return None
if node in hashd: return hashd[node]
node_copy = Node(node.val, [])
hashd[node] = node_copy
for n in node.neighbors:
n_copy = self.dfs(n, hashd)
if n_copy:
node_copy.neighbors.append(n_copy)
return node_copy
2-4) Bus Routes
# LC 815. Bus Routes
# V0
# IDEA : BFS + GRAPH
class Solution(object):
def numBusesToDestination(self, routes, S, T):
# edge case:
if S == T:
return 0
to_routes = collections.defaultdict(set)
for i, route in enumerate(routes):
for j in route:
to_routes[j].add(i)
bfs = [(S, 0)]
seen = set([S])
for stop, bus in bfs:
if stop == T:
return bus
for i in to_routes[stop]:
for j in routes[i]:
if j not in seen:
bfs.append((j, bus + 1))
seen.add(j)
routes[i] = [] # seen route
return -1
2-5) Course Schedule
// java
// V0
// IDEA : DFS (fix by gpt) (NOTE : there is also TOPOLOGICAL SORT solution)
// NOTE !!! instead of maintain status (0,1,2), below video offers a simpler approach
// -> e.g. use a set, recording the current visiting course, if ANY duplicated (already in set) course being met,
// -> means "cyclic", so return false directly
// https://www.youtube.com/watch?v=EgI5nU9etnU
public boolean canFinish(int numCourses, int[][] prerequisites) {
// Initialize adjacency list for storing prerequisites
/**
* NOTE !!!
*
* init prerequisites map
* {course : [prerequisites_array]}
* below init map with null array as first step
*/
Map<Integer, List<Integer>> preMap = new HashMap<>();
for (int i = 0; i < numCourses; i++) {
preMap.put(i, new ArrayList<>());
}
// Populate the adjacency list with prerequisites
/**
* NOTE !!!
*
* update prerequisites map
* {course : [prerequisites_array]}
* so we go through prerequisites,
* then append each course's prerequisites to preMap
*/
for (int[] pair : prerequisites) {
int crs = pair[0];
int pre = pair[1];
preMap.get(crs).add(pre);
}
/** NOTE !!!
*
* init below set for checking if there is "cyclic" case
*/
// Set for tracking courses during the current DFS path
Set<Integer> visiting = new HashSet<>();
// Recursive DFS function
for (int c = 0; c < numCourses; c++) {
if (!dfs(c, preMap, visiting)) {
return false;
}
}
return true;
}
private boolean dfs(int crs, Map<Integer, List<Integer>> preMap, Set<Integer> visiting) {
/** NOTE !!!
*
* if visiting contains current course,
* means there is a "cyclic",
* (e.g. : needs to take course a, then can take course b, and needs to take course b, then can take course a)
* so return false directly
*/
if (visiting.contains(crs)) {
return false;
}
/**
* NOTE !!!
*
* if such course has NO preRequisite,
* return true directly
*/
if (preMap.get(crs).isEmpty()) {
return true;
}
/**
* NOTE !!!
*
* add current course to set (Set<Integer> visiting)
*/
visiting.add(crs);
for (int pre : preMap.get(crs)) {
if (!dfs(pre, preMap, visiting)) {
return false;
}
}
/**
* NOTE !!!
*
* remove current course from set,
* since already finish visiting
*
* e.g. undo changes
*/
visiting.remove(crs);
preMap.get(crs).clear(); // Clear prerequisites as the course is confirmed to be processed
return true;
}
Decision Framework
Pattern Selection Strategy
Graph Algorithm Selection Flowchart:
1. What is the problem asking for?
├── Find shortest path → Continue to 2
├── Check connectivity → Use Union-Find or DFS/BFS
├── Order with dependencies → Use Topological Sort
├── Detect cycles → Use DFS with states or Union-Find
└── Traverse all nodes → Use BFS or DFS
2. For shortest path problems:
├── Unweighted graph → Use BFS
├── Non-negative weights → Use Dijkstra
├── Negative weights → Use Bellman-Ford
└── All pairs → Use Floyd-Warshall
3. For connectivity problems:
├── Static graph → Use DFS/BFS once
├── Dynamic connections → Use Union-Find
└── Count components → Use either approach
4. For traversal problems:
├── Level-by-level → Use BFS
├── Path finding → Use DFS with backtracking
└── State space search → Use BFS for optimal
5. Is the graph special?
├── Tree → Simpler DFS/BFS
├── DAG → Topological sort possible
├── Bipartite → Two-coloring
└── Grid → Treat as implicit graph
Algorithm Selection Guide
| Problem Type | Best Algorithm | Time | When to Use |
|---|---|---|---|
| Single-source shortest (unweighted) | BFS | O(V+E) | Simple shortest path |
| Single-source shortest (weighted) | Dijkstra | O((V+E)logV) | Non-negative weights |
| All-pairs shortest | Floyd-Warshall | O(V³) | Dense graphs |
| Cycle detection | DFS | O(V+E) | Directed graphs |
| Connected components | Union-Find | O(α(n)) | Dynamic connectivity |
| Topological order | Kahn’s/DFS | O(V+E) | Task scheduling |
| Minimum spanning tree | Kruskal/Prim | O(ElogE) | Network design |
Summary & Quick Reference
Complexity Quick Reference
| Algorithm | Time Complexity | Space Complexity | Notes |
|---|---|---|---|
| BFS/DFS | O(V + E) | O(V) | Standard traversal |
| Dijkstra | O((V+E)logV) | O(V) | With binary heap |
| Bellman-Ford | O(VE) | O(V) | Handles negative weights |
| Floyd-Warshall | O(V³) | O(V²) | All pairs |
| Union-Find | O(α(n)) | O(V) | Near constant |
| Topological Sort | O(V + E) | O(V) | Linear time |
Graph Building Patterns
Adjacency List
# For edges list
graph = defaultdict(list)
for u, v in edges:
graph[u].append(v)
graph[v].append(u) # Undirected
# For weighted edges
graph = defaultdict(list)
for u, v, w in edges:
graph[u].append((v, w))
Adjacency Matrix
# For unweighted
graph = [[0] * n for _ in range(n)]
for u, v in edges:
graph[u][v] = 1
graph[v][u] = 1 # Undirected
# For weighted
graph = [[float('inf')] * n for _ in range(n)]
for u, v, w in edges:
graph[u][v] = w
Common Patterns & Tricks
Visited Tracking
# Set for simple visited
visited = set()
# Array for state tracking
# 0: unvisited, 1: visiting, 2: visited
state = [0] * n
# Dictionary for path reconstruction
parent = {}
Grid as Graph
# 4-directional movement
directions = [(0, 1), (1, 0), (0, -1), (-1, 0)]
# 8-directional movement
directions = [(0, 1), (1, 0), (0, -1), (-1, 0),
(1, 1), (1, -1), (-1, 1), (-1, -1)]
# Check bounds
def is_valid(r, c, rows, cols):
return 0 <= r < rows and 0 <= c < cols
Cycle Detection Patterns
# Directed graph - DFS with states
def has_cycle_directed(graph):
# 0: unvisited, 1: visiting, 2: visited
state = [0] * n
def dfs(node):
if state[node] == 1: # Back edge
return True
if state[node] == 2:
return False
state[node] = 1
for neighbor in graph[node]:
if dfs(neighbor):
return True
state[node] = 2
return False
# Undirected graph - Union-Find
def has_cycle_undirected(edges):
uf = UnionFind(n)
for u, v in edges:
if not uf.union(u, v):
return True # Already connected
return False
Problem-Solving Steps
- Identify graph type: Directed/undirected, weighted/unweighted
- Choose representation: Adjacency list vs matrix
- Select algorithm: Based on problem requirements
- Handle edge cases: Empty graph, disconnected components
- Track state properly: Visited nodes, paths, distances
- Optimize if needed: Space or time improvements
Common Mistakes & Tips
🚫 Common Mistakes:
- Not handling disconnected components
- Incorrect visited state management
- Missing cycle detection in recursive DFS
- Wrong graph representation choice
- Not considering edge cases (self-loops, multiple edges)
✅ Best Practices:
- Use adjacency list for sparse graphs
- Clear visited tracking strategy
- Handle both directed and undirected cases
- Consider using Union-Find for dynamic connectivity
- Test with disconnected components
Interview Tips
- Clarify graph properties: Directed? Weighted? Connected?
- Draw small examples: Visualize the problem
- Choose right representation: List vs matrix
- State complexities: Time and space upfront
- Handle edge cases: Empty, single node, cycles
- Optimize incrementally: Start simple, then improve
Related Topics
- Trees: Special case of graphs (connected, acyclic)
- Dynamic Programming: DP on graphs (paths, trees)
- Greedy Algorithms: MST algorithms
- Heap/Priority Queue: Used in Dijkstra, Prim’s
- Recursion/Backtracking: DFS implementation
2-6) Find Eventual Safe States
// java
// LC 802
// V1-0
// IDEA : DFS
// KEY : check if there is a "cycle" on a node
// https://www.youtube.com/watch?v=v5Ni_3bHjzk
// https://zxi.mytechroad.com/blog/graph/leetcode-802-find-eventual-safe-states/
public List<Integer> eventualSafeNodes(int[][] graph) {
// init
int n = graph.length;
State[] states = new State[n];
for (int i = 0; i < n; i++) {
states[i] = State.UNKNOWN;
}
List<Integer> result = new ArrayList<>();
for (int i = 0; i < n; i++) {
// if node is with SAFE state, add to result
if (dfs(graph, i, states) == State.SAFE) {
result.add(i);
}
}
return result;
}
private enum State {
UNKNOWN, VISITING, SAFE, UNSAFE
}
private State dfs(int[][] graph, int node, State[] states) {
/**
* NOTE !!!
* if a node with "VISITING" state,
* but is visited again (within the other iteration)
* -> there must be a cycle
* -> this node is UNSAFE
*/
if (states[node] == State.VISITING) {
return states[node] = State.UNSAFE;
}
/**
* NOTE !!!
* if a node is not with "UNKNOWN" state,
* -> update its state
*/
if (states[node] != State.UNKNOWN) {
return states[node];
}
/**
* NOTE !!!
* update node state as VISITING
*/
states[node] = State.VISITING;
for (int next : graph[node]) {
/**
* NOTE !!!
* for every sub node, if any one them
* has UNSAFE state,
* -> set and return node state as UNSAFE directly
*/
if (dfs(graph, next, states) == State.UNSAFE) {
return states[node] = State.UNSAFE;
}
}
/**
* NOTE !!!
* if can pass all above checks
* -> this is node has SAFE state
*/
return states[node] = State.SAFE;
}